MicroRNA and liver cancer
Abstract
Hepatocellular carcinoma (HCC) is a major cause of cancer-related deaths worldwide. HCC is characterized by a poor prognosis and an ever increasing number of scientific studies aim to find new diagnostic, prognostic, and therapeutic targets. MicroRNAs (miRNAs), small non-coding RNAs that regulate the gene expression in many processes, have been shown to play a crucial role in regulating hepatocellular carcinoma. miRNAs may act as oncogenic miRNAs and tumor suppressor miRNAs and regulate cancer cell proliferation, invasion, and metastasis by being differently upregulated or downregulated and targeting the genes related with carcinogenesis. miRNAs secreted from cancer cells are found circulating in the blood, presenting an opportunity for their use as disease-related biomarkers. Moreover, extracellular vesicle-derived miRNAs are known to reflect the cell of origin and function and may provide effective biomarkers for predicting diagnosis and prognosis and new therapeutic target in HCC. In this article, we describe the most recent findings regarding the molecular mechanisms and gene regulation of microRNA in HCC, as well as their application in diagnosis/prognosis and treatment.
Keywords
Introduction
Hepatocellular carcinoma (HCC) is considered the second leading cause of cancer-related death worldwide[1]. Although several treatment options, such as surgical resection, liver transplantation, percutaneous ablation, and administration of anticancer drugs such as solafenib, are available, a high recurrence rate and unsatisfactory survival rate are major obstacles to the treatment of HCC[2]. Because chronic infection by hepatitis viruses, especially hepatitis C virus (HCV), is a main cause of HCC, the application of revolutionary, highly effective direct-acting antiviral agents against HCV, which have recently become available, is a promising approach to reducing future HCC incidence. However, the considerable risk of HCC development even after eradication of HCV remains a challenge[3]. In addition, hepatitis B virus (HBV)-related as well as alcohol/non-alcoholic liver disease-related HCC is also an important public health burden. Although extensive efforts have identified several driver genes frequently mutated in HCC[4,5], potentially curative treatments are limited. New drugs (i.e., lenvatinib, regorafenib, cabozantinib, and ramucirumab) have been shown to improve the clinical outcomes of patients with advanced HCC but improvements in terms of survival are modest.
MicroRNAs (miRNAs) are small, non-coding RNAs of approximately 21-23 nucleotides and are broadly conserved in eukaryotes, including plants and animals. miRNAs downregulate the expression of their target genes post-transcriptionally by complementary base pairing with the 3’-untranslated region of messenger RNAs (mRNAs) and, in doing so, control various fundamental biological processes[6]. In the last decade, pivotal roles of miRNAs in the development and progression of cancer have become widely been accepted[7]. Extensive efforts have shown that the expression profiles of miRNAs are frequently altered in HCC, and miRNAs have been shown to contribute to the development and progression of HCC. In this review, we summarize recent progress in the field of HCC-related miRNAs, application of miRNAs to the diagnosis of HCC, and their therapeutic potential for the treatment of HCC.
miRNA biogenesis and function
miRNAs are encoded in the genome either as independent gene segments or within the introns of protein-coding genes. miRNA-encoding genes are first transcribed by RNA polymerase II, generating long transcripts called “primary miRNAs” that contain several stem-loop secondary structures[8]. After capping at the 5’ end and polyadenylation at the 3’ end, primary miRNAs are processed in the nucleus by the endonuclease Drosha and cleaved into precursor miRNAs, which comprise a mature miRNA sequence and a complementary sequence linked by a short loop region[9]. After being transported from the nucleus to the cytoplasm by exprotin-5, the precursor miRNA stem-loop is further processed by RNase III endonuclease Dicer to produce an RNA duplex. The RNA duplex is incorporated into the RNA-induced silencing complex (RISC), which carries out miRNA-mediated post transcriptional regulation of the target genes. Basically, only one strand of the RNA duplex is utilized as a mature miRNA and the opposite complementary strand (miRNA*) is removed from the RISC complex[10]. A 6-7 nt sequence complementary to the 5’ end of each miRNA is present in the 3’ untranslated region or, in some cases, other regions of the target mRNAs. The Cyclin D1 (CCND1), Mesenchymal-epithelial transition factor (cMET) and CDK6 suppresses translation by binding to the target mRNA. Because each miRNA can recognize hundreds of target genes and each protein coding gene is controlled by several miRNAs, miRNAs can broadly control complex gene regulatory networks. In addition, recent studies have suggested that some miRNAs contribute to the stability and robustness of gene regulatory networks, which are influenced by environmental or intrinsic fluctuations, such as temperature and genetic/epigenetic variations[11]. Collectively, miRNAs serve as fine-tuners for the precise control of various cellular responses.
Role of miRNA in HCC
In this paper, we introduce the impact of miRNAs on HCC growth, progression, and tumor microenvironment. Several large-scale analyses of miRNA expression profile for HCC provided the key miRNAs. miRNA has many putative miRNA recognition sites, and there are many target genes predicted by computational tools. It is difficult to fully understand their functions, but recent studies have gradually elucidated the role of miRNAs in HCC. Moreover, recent evidence suggests that the epigenetic mechanisms of miRNA expression provide a potential therapeutic target[11]. Research of not only the expression but also the methylation of miRNAs may be important to better understand the relationship between miRNA and HCC.
miRNA expression profiles in HCC
Several studies have been conducted to search for biomarkers or therapeutic agents by investigating the differential expression of miRNAs between hepatocellular carcinoma tissue and the corresponding non-tumor liver tissue.
Andrés-León et al.[12] analyzed miRNAs in The Cancer Genome Atlas (TCGA) data for HCC and other tumor types. In 2018, a study integrated miRNA expression database (TCGA, GSE31384[13], and GSE6857[14]) and compared their expression in liver cancer and adjacent normal samples to investigate the difference of miRNAs expression associated with carcinogenesis[15]. They found that hsa-miR-149, hsa-miR-139, hsa-miR-3677, hsa-miR-550a, and hsa-miR-212 were significantly correlated with overall survival of HCC patients. Li et al.[16] also undertook the integrated analysis of a number of miRNA profiling studies[17-34] in HCC samples to detect differentially expressed miRNAs. As the result of analysis, miR-221/222, miR-195, and miR-199a were consistently differentially expressed in multiple independent studies.
Although no overlapping expression patterns of some miRNAs could be found in the existing literature and the currently available data, the miRNAs reported most consistently as dysregulated, namely hsa-miR-18, hsa-miR-21, hsa-miR-106, hsa-miR-221/222, hsa-miR-224, hsa-miR-99, hsa-miR-195, and hsa-miR-199, play a role in regulating the hallmarks of HCC [Table 1].
Summary of consistently reported up-/downregulated microRNAs in profiling studies
| Upregulated in HCC tissues | Ref. |
|---|---|
| hsa-miR-18a-3p | [12,16,17] |
| hsa-miR-21-5p | [12,15-19] |
| hsa-miR-25-3p | [15,19] |
| hsa-miR-93-5p | [12,18,19] |
| hsa-miR-96-5p | [12,15] |
| hsa-miR-106b-5p | [15,17,19] |
| hsa-miR-151-3p | [17,19] |
| hsa-miR-151a-5p | [12,17] |
| hsa-miR-181b-5p | [12,15] |
| hsa-miR-182-5p | [12,15] |
| hsa-miR-183-3p | [12,15] |
| hsa-miR-183-5p | [12,15] |
| hsa-miR-210 | [16,17] |
| hsa-miR-221 | [12,15-17] |
| hsa-miR-222 | [12,15-18] |
| hsa-miR-224-5p | [12,16-18] |
| hsa-miR-301b | [12,15] |
| Downregulated in HCC tissues | |
| hsa-let-7c-5p | [12,19] |
| hsa-miR-99a-5p | [16,17,19] |
| hsa-miR-125b | [16,17] |
| hsa-miR-130a-3p | [12,18] |
| hsa-miR-139-3p | [12,15] |
| hsa-miR-139-5p | [12,15,19] |
| hsa-miR-142-5p | [12,17] |
| hsa-miR-145 | [12,15,18] |
| hsa-miR-195 | [12,15-18] |
| hsa-miR-199a-3p | [15,17-19] |
| hsa-miR-199a-5p | [16,18,19] |
| hsa-miR-214-3p | [12,15,18] |
| hsa-miR-326 | [12,15] |
| hsa-miR-424-3p | [12,15] |
| hsa-miR-424-5p | [12,15] |
miRNA as tumor suppressors
A tumor suppressor gene is defined as a gene which encodes proteins that inhibit tumorigenesis. Loss or reduction of function in tumor suppressor genes plays an oncogenic role, leading to the development of many tumors. The product of the tumor suppressor genes plays a regulatory role of cell proliferation, cell differentiation and DNA repair. Notably, as shown in Table 1, several miRNAs are significantly downregulated in tumor tissues and may be the characteristics of tumor suppressor genes.
Recent studies have revealed that miR-199 inhibited HCC proliferation and metastasis. miR-199a/b-3p can suppress HCC growth in vitro and in vivo by inhibiting inhibiting p21-Activated kinase 4 (PAK4), which is known to activate Raf/MEK/ERK pathway[35]. Moreover, in another study, miR-199a-5p was shown to reprogram HCC cell glycolysis by directly targeting HK2 (hexokinase 2) in HCC[36]. miR-195 regulated cell cycle progression via targeting CDK6, CCNE1, CDC25A, and CDK4, leading to the abnormal cell proliferation in HCC development[37]. miR-122 is known to be a highly expressed miRNA and one of the most abundant liver specific miRNAs, accounting for about 70% and 52% of the whole hepatic total miRNAs in adult mice and humans, respectively[38-40]. miR-122 also plays a significant role in the regulation of cholesterol and fatty acid metabolic pathway in the liver, and reduction of miR-122 expression in the liver contributes to the development of steatohepatitis[41-43]. Tsai et al.[44] reported that liver-specific miR-122 knockout mice appear normal but develop steatohepatitis for dysregulation of lipid metabolism. Importantly, knockdown of miR-122 in the liver also promoted development of liver fibrosis and carcinogenesis. In addition, Simerzin et al.[45] characterized one mechanism by which reduction of miR-122 expression induces HCC, and revealed that miR-122 activated p53 by targeting Mdm2 (mouse double minute 2 homolog), known as the negative regulator of p53. Roy et al.[46] reported integrated analysis of the miRNA and mRNA expression in the liver tissues from a mouse model of HCC and observed a reduction of miR-193a-5p in murine and human HCC cells and tissues, leading to increased levels of NUSAP1 (nucleolar- and spindle-associated protein), which regulates cell proliferation by controlling spindle assembly and genomic stability[47]. In fact, an miR-193a-5p mimic and knockdown of NUSAP1 in Huh7 cells suppressed HCC development and migration. miR-206 inhibits tumor growth in HCC by targeting CCND1, cMET, and CDK6, and delivery of miR-206 into the liver almost completely suppressed growth of HCC in an AKT/Ras and cMyc HCC mouse model[48].
The Wnt/β-catenin signaling pathway plays an essential role in the active process of liver development and governs the proliferation and differentiation of hepatocytes. Abnormal activity and gene mutations of this pathway are known to be involved in HCC development[49]. miR-148b targeting WNT1 contributes to cell growth in HCC and functions as a tumor suppressor in HCC[50]. Another study suggested that downregulation of miR-766-3p promotes HCC cell progression by targeting the Wnt3/ Protein regulator of cytokinesis-1 pathway as a tumor suppressor gene in HCC[51].
miRNAs as oncogenes
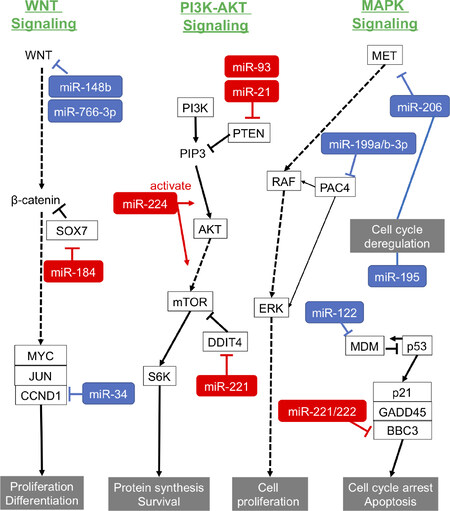
Recent studies have shown that some miRNAs, called oncogenic miRNAs (oncomiRs), regulate cell proliferation and the apoptotic processes that are important in HCC forming and growth [Table 2 and Figure 1].
List of oncomiRs and suppressor miRNAs
| OncomiRs | Target(s) | Ref. |
|---|---|---|
| miR-21 | PTEN | [52] |
| HBP1-p53-SREBP1C pathway | [53] | |
| miR-93 | PTEN, CDKN1A | [60] |
| miR-106b | DAB2 (disabled homolog 2) | [61] |
| miR-184 | SOX7 | [59] |
| miR-221 | p53 | [62] |
| p27 and/or DDIT4 | [56] | |
| miR-221/222 cluster | BBC3 | [57] |
| miR-224 | AKT signaling | [58] |
| Tumor suppressor miRNAs | ||
| miR-122 | p53/MDM2 | [45] |
| miR-148b | WNT1 | [50] |
| miR-193a-5p | NUSAP1 | [46] |
| miR-195 | CDK6, CCNE1, CDC25A, and CDK4 | [37] |
| miR-199a/b-3p | PAK4/Raf/MEK/ERK pathway | [35] |
| miR-199a-5p | HK2 | [36] |
| miR-206 | CCND1, cMET, and CDK6 | [48] |
| miR-766-3p | Wnt3a/PRC1 pathway | [51] |
Figure 1. Roles of oncomiRs and suppressor miRNAs in carcinogenesis pathways of HCC. Red: oncomiRs; blue: suppressor miRNAs. PTEN: phosphatase and tensin homolog; oncomiRs: oncogenic miRNAs; miRNAs: microRNAs; HCC: hepatocellular carcinoma; PI3K: phosphoinositide 3-kinase; MAPK: mitogen-activated protein kinases; MET: mesenchymal-epithelial transition factor; PAC4: proteasome assembly chaperone 4 ; ERK: extracellular regulated kinase; MDM: mouse double minute; BBC3: Bcl-2-binding component 3; GADD45: growth arrest and DNA-damage-inducible 45 alpha; S6K: ribosomal protein S6 kinase; DDIT4: DNA-damage-inducible transcript 4; mTOR: mechanistic target of rapamycin; SOX7: SRY-box 7; CCND1: Cyclin D1
Phosphatase and tensin homolog (PTEN) is a tumor suppressor gene and is involved in the regulation of the cell cycle, preventing cells from growing and dividing rapidly. Excessive expression of miR-21 can promote HCC growth and spread by inhibiting PTEN expression[52]. Moreover, another study revealed that miR-21 contributes to hepatic steatosis and cancer progression by modulating p53 and Srebp1c pathway via target HBP1 (HMG-box transcription factor 1)[53]. miR-221 is over-expressed and associated with the tumor capsular infiltration that affects invasion and metastasis in HCC clinical samples, and miR-221 inhibits cell proliferation and apoptosis-mediating G1/S-phase arrest[54]. Pineau et al.[55] showed that miR-221 overexpression promoted the growth of tumorigenic murine hepatic progenitor cells using a mouse HCC model, and that miR-221 could induce the tumor growth via suppressing p27 and/or DDIT4 (DNA damage-inducible transcript 4) expression. miR-222, which is in the miR-221/222 cluster and is upregulated in HCC tissues, promotes the progression of HCC by targeting the 3’UTR of the mRNA of BBC3 (Bcl-2 binding component 3)[56].
Ma et al.[57] showed that miR-224 acted as an oncogenetic regulator of HCC through the activation of AKT signaling[58]. miR-184 upregulated c-Myc and cyclin D1 and phosphorylated Rb protein by targeting SOX7, resulting in increasing proliferation in HCC cells[59]. As shown in Table 2, the targets of miR-93, miR-106b and miR-221 were PTEN, CDKN1A[60], DAB2[61] and p53[62], respectively.
Recent findings suggest that miRNAs are involved in the abnormal epigenetic regulation of cancer cells[63]. In HCC, a recent study showed hypomethylation of a cancer-specific microRNA cluster in tumors from patients without HCV or HBV infection[64]. He et al.[65] determined that oncomiRs, including miR-106b, miR-25, miR-93, miR-23a, and miR-27a, are hypomethylated and upregulated in HCC. Moreover, methylation of mature miRNAs may be related with carcinogenesis. Konno et al.[66] showed that methylated miRNAs were increased in gastrointestinal cancer and methylated miR-200c-3p suppressed its target genes compared with unmethylated miR-200c-3p. In HCC, methylated miRNAs have not been reported yet, but it may be necessary to examine the relationship between methylation of miRNAs and tumors as well as expression of miRNAs.
Involvement of miRNAs in HCC invasion and metastasis
Recent studies suggest the aberrant expression of miRNAs also affects the invasion and metastasis of HCC. miR-21, miR-34a, and miR-224 have been shown to be involved in proliferation, epithelial-mesenchymal transition (EMT), and metastasis of HCC[67,68].
Zhou et al.[69] reported that tumor-derived extracellular vesicles (EVs) miR-21 could convert hepatic stellate cells to cancer-associated fibroblasts by decreasing PTEN expression, leading to activation of the PDK1/AKT signaling pathway in HCC. In another study, miR-21 produced by endothelial progenitor cells led to cell migration, invasion, and EMT via induction of monocyte chemotactic protein 1 in liver cancer cells[70]. miR-224 inhibited Homeobox D10 directly and activated expression of p-PAK4 and MMP-9, which regulated remodeling, invasion, and metastasis in tumors[71].
Extracellular vesicles are the endosome-derived vesicles with a diameter of 40-100 nm that are actively secreted by most cells, and they contain functional proteins, mRNAs, and miRNAs. It has been elucidated that HCC cell-secreted EVs can transfer miRNAs into recipient cancer cells, tumor microenvironment, and distant organs, thus promoting growth, migration, and invasion[72,73]. Lin et al.[74] reported that EV-derived miR-210 secreted by HCC cells could be delivered into endothelial cells and directly suppressed SMAD4 and STAT6 expression, thereby promoting tumor angiogenesis. Hepatoma cell-secreted EV-derived miR-103 increases vascular permeability and enhanced tumor metastasis by inhibiting multiple endothelial junction proteins [VE-Cadherin (VE-Cad), p120-catenin (p120), and tight junction][75]. Fang et al.[76] showed that EV-derived miR-1247-3p secreted by highly metastatic HCC cells activates β1-integrin-NF-κB signaling in fibroblasts and converts into cancer-associated fibroblasts, resulting in the promotion of cancer metastasis.
miRNAs and tumor immunity
In a recent study, immune checkpoint therapy targeting T cell-negative costimulatory molecules such as programmed cell death-1(PD-1) showed a therapeutic effect at a certain rate in many cancers. Emerging evidence suggests that tumor-derived microRNAs involve tumor immune escape by targeting immune checkpoint proteins and molecules[77]. Several miRNAs have been found to be downregulated in cancer cells to allow PD-L1 expression[78,79].
In HCC, cancer-derived EVs also suppress CD8+ T-cell function by promoting Bregs, Tregs, or M2 macrophages[80]. HCC-derived miR-146a upregulated PD-1, TIGIT, and CTLA4 on T cells by promoting M2 tumor-associated macrophages and accelerated HCC progression[81]. Secreted EVs miR-23a from HCC cells under endoplasmic reticulum-stress conditions increased PD-L1 expression on macrophages, which then decreased expansion and function and promoted apoptosis in cytotoxic T lymphocyte[82].
These miRNAs can be predictors of tumor response to treatment but also might be new therapeutic targets for HCC treatment. There is still a lack of research in this field, and further investigation of the relationship between tumor immunity and miRNAs is expected.
Application of circulating miRNAs as clinical biomarkers
Early diagnosis is beneficial for the prognosis of cancer, including HCC. Serum alpha fetoprotein (AFP) is the most used biomarker for diagnosis of HCC but its sensitivity and specificity are only 60% and 90%, respectively[83]. Reliable and simple prognostic factors have not been established yet, but the identification of circulating miRNAs that are useful as biomarkers for HCC diagnosis, prognosis, and therapeutic response is expected[84,85].
Circulating miRNA as diagnostic biomarkers for HCC
Several reports have shown the potential of circulating miRNAs as a clinical biomarker in HCC diagnosis [Table 3].
Circulating microRNAs as diagnostic biomarkers for HCC
| Ref. | Body fluid | Case | Number | Control/Number | microRNAs | Regulation |
|---|---|---|---|---|---|---|
| Zhou et al.[86] | Plasma | HCC | 457 | NH167/CHB169/LC141 | miR-21 | UP |
| miR-192 | UP | |||||
| miR-801 | UP | |||||
| miR-26a | DN | |||||
| miR-27a | DN | |||||
| miR-122 | DN | |||||
| miR-223 | DN | |||||
| Jin et al.[87] | Plasma | HCC | 116 | NH79/CHB49/LC18 | miR-1972 | UP |
| miR-193a-5p | UP | |||||
| miR-214-3p | UP | |||||
| miR-365a-3p | UP | |||||
| Tan et al.[88] | Serum | HCC | 261 | NH173/LC233 | miR-206 | UP |
| miR-141-3p | UP | |||||
| miR-433-3p | UP | |||||
| miR-1228-5p | UP | |||||
| miR-199a-5p | DN | |||||
| miR-122-5p | DN | |||||
| miR-192-5p | DN | |||||
| miR-26a-5p | DN | |||||
| Zhang et al.[89] | Serum | HCC | 115 | NH40 | miR-16-2-3p | UP |
| miR-92a-3p | UP | |||||
| miR-107 | UP | |||||
| miR-3126-5p | DN | |||||
| Huang et al.[90] | HCC | 3423 | NH1887/CHB2403 | miR-21 | UP | |
| miR-122 | UP | |||||
| Yamamoto et al.[91] | Serum | HCC | 353 | NH1033/CH46/LC93 | miR-320b | UP |
| miR-663a | UP | |||||
| miR-4448 | UP | |||||
| miR-4651 | UP | |||||
| miR-4749-5p | UP | |||||
| miR-6724-5p | UP | |||||
| miR-6877-5p | UP | |||||
| miR-6885-5p | UP |
Zhou et al.[86] investigated plasma microRNA expression in three independent cohorts, comprising 934 participants (healthy, chronic hepatitis B, cirrhosis, and HBV-related HCC). A microRNA panel (miR-122, miR-192, miR-21, miR-223, miR-26a, miR-27a, and miR-801) revealed high sensitivity of HCC diagnosis (AUC = 0.864 and 0.888 for the training and validation datasets, respectively). Jin et al.[87] reported the plasma circulating miRNA profiling of 262 samples (116 HCC patients, 18 cirrhosis patients, 49 chronic hepatitis B, and 79 healthy controls) from three independent cohorts to find effective biomarkers that can distinguish the different liver disease stages of chronic hepatitis, cirrhosis, and HCC. They found four miRNAs (miR-1972, miR-193a-5p, miR-214-3p, and miR-365a-3p) that could distinguish HCC from non-HCC samples.
Tan et al.[88] conducted a large-scale study of 667 subjects (261 HCC patients, 233 cirrhosis patients, and 173 healthy controls) to identify serum circulating miRNAs as reliable biomarkers in the diagnosis of HBV-related HCC. They identified an miRNA set (hsa-miR-206, hsa-miR-141-3p, hsa-miR-433-3p, hsa-miR-1228-5p, hsa-miR-199a-5p, hsa-miR-122-5p, hsa-miR-192-5p, and hsa-miR-26a-5p) that provided high diagnostic accuracy for HCC (healthy: AUC = 0.893; cirrhosis: AUC = 0.892). Zhang et al.[89] also identified a miRNAs panel (miR-92-3p, miR-107, and miR-3126-5p) as a reliable diagnostic marker for HCC, especially for early stage patients (AUC = 0.975) and for low-level AFP HCC patients (AUC = 0.971). Huang et al.[90] performed comprehensive analysis of large Chinese datasets in the Medline, Embase, and Chinese National Knowledge Infrastructure (MEDLINE) databases to elucidate effective miRNAs for the early diagnosis of HCC. miR-21 and miR-122 were significantly upregulated in multiple studies on HCC and capable of diagnosing HCC. However, since this result may be specific to ethnic groups, an analysis that integrates multiracial data will be required in the future. Recently, Yamamoto et al.[91] reported that an eight-miRNA panel (miR-320b, miR-663a, miR-4448, miR-4651, miR-4749-5p, miR-6724-5p, miR-6877-5p, and miR-6885-5p) could accurately distinguish HCC patients from healthy (AUC = 1.00; sensitivity, 97.7%; and specificity, 98.4%) and chronic hepatitis/cirrhosis patients (AUC = 0.99; sensitivity, 97.7%; and specificity, 94.7%) in a study of 344 HCC patients. Importantly, compared with serum AFP, the eight-miRNA panel showed high sensibility (stage I, 98%) at the early stage of HCC. Furthermore, they have started the prospective analysis of the eight-miRNA panel for HCC screening.
Validating and quantifying of miRNA can be influenced by the type of blood sample, collection protocol, and isolation and detection methods. The lack of standardized detection protocols and data normalization makes it difficult to compare the results of different studies, therefore leading to inconsistent findings in screening of circulating miRNAs. Recently, in a comprehensive analysis of miRNA expression in serum and plasma, there are distinct differences in both the number and quantification of detected miRNAs between serum and plasma samples[92]. Thus, it is necessary to use standardized protocols, including for sample collection, RNA isolation, and the selection of a suitable internal control for normalization.
Circulating miRNA as prognostic and recurrence markers for HCC
Xu et al.[93] analyzed the correlation between serum miR-122 expression and prognosis in 122 HCC patients, and their results showed that HCC patients with high expression of serum miR-122 had significantly better overall survival rate than those with low miR-122 levels. Over-expression of pretreatment plasma miR-122 is also correlated with early refractoriness in HCC patients with transarterial chemoembolization (TACE) treatment[94,95]. Li et al.[96] found that high levels of serum miR-221 expression was associated with tumor size, cirrhosis, and tumor stage in 46 HCC patients. Their results suggest that circulating miR-221 expression could be a prognostic marker. In a large screening study, Zhang et al.[97] investigated the potential prognostic miRNAs using data from 54 relevant articles and 6464 patients and identified circulating miR-148a and miR-192 as reliable prognostic markers in HCC. Jin et al.[87] also revealed that six miRNAs (miR-424-5p, miR-101-3p, miR-128, miR-139-5p, miR-382-5p, and miR-410) were significantly related with overall survival rate in 116 HCC patients and had potential to be good biomarkers for predicting prognosis.
Several studies identified circulating miRNAs as therapy predictive markers[98]. Chuma et al.[99] reported that serum miR-1246 predicted early tumor recurrence after hepatic resection. Cho et al.[100] revealed that the levels of circulating miR-26a and miR-29a before treatment were prognostic markers for liver transplantation-free survival in hepatitis B virus-related HCC patients. Serum levels of miR-181a-5p and miR-122 were found to have potential to predict response of HCC to sorafenib[101,102]. Recently. Teufel et al.[103] reported some plasma miRNAs associated with response to regorafenib treatment in HCC.
Extracellular vesicle-derived miRNAs in diagnosis
miRNAs are more stable in serum exosomes because the lipid bilayer protects them from degradation by RNases[104]. Therefore, blood samples are appropriate for the evaluation of extracellular vesicle-derived miRNAs as potential biomarkers in HCC diagnosis and prognosis. Moreover, EVs can specifically reflect their original cell types and conditions and may be able to detect early stages of cancer as special biomarkers[105-107].
In their recent studies, Sohn et al.[108] isolated EVs from the serum of CHB, cirrhosis, and HCC patients and found over-expression of several EV-derived miRNAs, namely miR-18a, miR-221, miR-222, and miR-224, and downregulation of serum EV-derived miR-101, miR-106b, miR-122, and miR-195 in HCC patients. Additionally, they compared expression patterns of EV-derived miRNAs with those of serum circulating miRNAs. They revealed that EV-derived miRNAs were more sensitive to distinguishing HCC from CHB/cirrhosis than serum circulating miRNAs. Other studies found high levels of EV-derived miR-21, an abundant miRNA expressed in HCC tissue, and this was an independent predictor of mortality and disease progression[109-111]. Mjelle et al.[112] analyzed paired microRNAs profiling of tumor and normal tissue and circulating EVs from HCC patients using small RNA sequencing methods. There was a consistent positive correlation of miR-21 expression between circulating EVs and HCC tissue.
To identify specific microRNAs in exosomes from the sera of patients with recurrent HCC, Sugimachi et al.[113] performed miRNA analyses of samples from 59 patients who underwent living donor liver transplantation. They found significantly lower levels of miR-718 in the serum EVs of HCC cases with recurrence. Downregulation of miR-718 increased expression of HOXB8, a known oncogene, resulting in carcinogenesis and metastasis. Another study showed serum EV-derived miR-122 and miR-21 could be predictive biomarkers in HCC patients with liver cirrhosis treated with TACE[114]. The research revealed that patients with a higher miR-122 ratio (after TACE/before TACE) had significantly longer disease-specific survival, compared to patients with a lower miR-122 ratio.
As mentioned above, although several studies have investigated the clinical significance of EV-derived miRNAs in HCC, there are still few reports on serum EV-derived microRNA in HCC. A larger cohort of HCC should be included and further improvements in the isolation and detection of serum EV-derived miRNA are expected.
Recently, lncRNAs and proteins in exosomes have also been the focus of research to understand the pathogenesis of cancer and its progression. Yuan et al.[115] showed that lncRNA-ATB acted as a key regulator of TGF-β signaling pathways and induced EMT and tumor cell invasion in HCC. Interestingly, Hoshino et al.[116] found EVs from different tumor types bear integrins that target these exosomes to specific organs and trigger signaling pathways, thereby initiating pre-metastatic niche formation. They suggested that the integrin contained in exosomes could be used as a biomarker to predict metastasis destination in cancer patients. The research of exosomes in HCC will contribute to gain new biological insights and provide novel diagnostics and therapeutics targets.
miRNA-based therapeutics in HCC
miRNAs are also considered as potential therapeutic targets for cancer. miRNA therapeutics are based on targeting oncomiRs or mimicking tumor suppressor miRNAs[117]. Double-stranded miRNA mimics aim to upregulate tumor suppressor miRNA, whereas anti-oncomiRs, short interfering RNA (siRNA), and single stranded anti-sense oligonucleotides are designed to specifically bind to and inhibit miRNAs that bear oncogenic properties.
Since there are abundant ribonucleases, administered miRNA is rapidly degraded in blood. One solution to this is chemical modification, e.g., using phosphorothioate-containing oligonucleotides, 2-O-methyl-(2-O-Me)/2-O-methoxyethyl oligonucleotides (2-O-MOE), 2-fluoro oligonucleotides (2’-F), and locked nucleic acid (LNA) oligonucleotides[118]. In liver disease, miravirsen, an LNA-containing anti-miR-122 that inhibits translation of HCV genome, entered phase II clinical trials in 2017, and the result of the clinical trial was positive[119-122]. N-acetylgalactosamine (GalNAc) platform enables specific, targeted uptake into hepatocytes. GalNAc-conjugated anti-miR targeting miR-103 and miR-107 that regulate insulin sensitivity is being used in patients with non-alcoholic fatty liver disease and type 2 diabetes in a Phase I/II trial[123,124].
Another solution for miRNA-based therapeutics is a drug delivery system. There are two types of miRNA delivery systems, viral and non-viral[125]. Viral vector systems usually use retroviruses, lentiviruses, and adenoviruses or adeno-associated viruses. Viral vectors have higher transfection efficiency but are more toxic and immunogenic. For this reason, safer non-viral carriers are being developed for clinical applications. Non-viral vectors include lipid-based nanocarriers, polymeric vectors/dendrimer-based vectors, and cell-derived membrane vesicles. A liposome-encapsulated miR-34a mimic (MRX34) was able to block HCC growth in more than a third of the treated animals[126]. Furthermore, MRX34 was used for patients with various tumors, including HCC (n = 14) in a Phase I trial. One patient with HCC achieved a prolonged PR lasting 48 weeks, and four patients experienced SD lasting more than four cycles[127]. Sato et al.[128] developed a multifunctional envelope nanodevice (MEND) incorporating various functions, including polyethylene glycol modification and introduction of membrane-permeable peptides. A pH-sensitive lipid YSK05 (YSK05-MEND) specifically delivered higher amounts of the anti-miR-122 to the liver[129]. Recently, extracellular vesicles are considered to play important roles in intercellular communication and are also used as effective drug carriers[130-132]. EVs have low cytotoxicity and antigenicity. Application studies on nanocarriers that encapsulate miRNA, siRNA, etc. inside extracellular vesicles by ultrasonic irradiation or electroporation and deliver them to target cells are being performed[133]. In a recent study, EVs were isolated from human plasma and engineered with miR-31 and miR-451a. The engineered EVs promoted apoptosis of HCC in vitro[134]. Moreover, in another study, EVs were isolated from human adipose tissue-derived mesenchymal stromal cells (ASC) modified with a lentiviral vector expressing miR-125b. The miR-125b containing EVs inhibited proliferation in HCC cells by suppressing p53 expression[135].
Conclusion
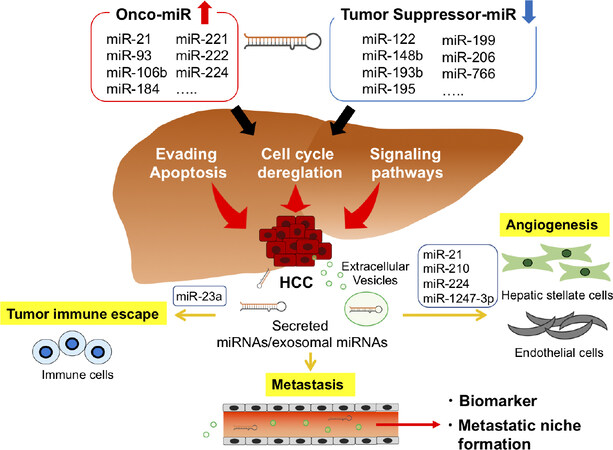
miRNAs are being considered as new biomarkers and potential therapeutic targets for HCC [Figure 2]. To date, many miRNAs have been identified as regulators of target genes involved in carcinogenesis, invasion, and metastasis in HCC. In addition, circulating EV-derived miRNAs secreted by cancer cells may become strong cancer biomarkers and novel therapeutic targets, and miRNA-based therapies have evolved and are being developed to be safer and more effective. Further research is needed to determine the prospects of new miRNA-based anticancer therapies for HCC treatment. However, several issues need to be resolved to adopt circulating miRNAs for diagnostic application in clinical routine. Normalization is one of the most controversial issues. The choice of a reference gene can have a significant impact on measuring the level of transcripts and, thus, on the biological interpretation of the data. In addition, standardization of miRNA processing is required, from sample collection and sample storage to RNA isolation and reverse-transcription. If these current limitations can be overcome, miRNAs may eventually be implemented in diagnostic algorithms, as well as be used to predict the clinical outcomes of HCC patients.
Declarations
Authors’ contributionsConcepted: Onishi M, Ochiya T, Tanaka Y
Designed: Onishi M, Tanaka Y
Wrote (first draft): Onishi M
Wrote, reviewed, discussed, edited, and revised: Onishi M, Ochiya T, Tanaka Y
Availability of data and materialsNot applicable.
Financial support and sponsorshipThis work was supported by Japan Agency for Medical Research and Development (AMED) under Grant Number JP18fk0210001 and JP19fk0210048, and JSPS KAKENHI under Grant Number JP19H03640.
Conflicts of interestAll authors declared that there are no conflicts of interest.
Ethical approval and consent to participateNot applicable.
Consent for publicationNot applicable.
Copyright© The Author(s) 2020.
REFERENCES
1. Fitzmaurice C, Abate D, Abbasi N, Abbastabar H, Abd-Allah F, et al; Global Burden of Disease Cancer Collaboration. Global, regional, and national cancer incidence, mortality, years of life lost, years lived with disability, and disability-adjusted life-years for 29 cancer groups, 1990 to 2017: a systematic analysis for the global burden of disease study. JAMA Oncol 2019;5:1749-68.
2. Xia F, Wu LL, Lau WY, Huan HB, Wen XD, et al. Adjuvant sorafenib after heptectomy for barcelona clinic liver cancer-stage C hepatocellular carcinoma patients. World J Gastroenterol 2016;22:5384-92.
3. Matsuura K, Sawai H, Ikeo K, Ogawa S, Iio E, et al. Genome-wide association study identifies TLL1 variant associated with development of hepatocellular carcinoma after eradication of hepatitis C virus infection. Gastroenterology 2017;152:1383-94.
4. Cleary SP, Jeck WR, Zhao X, Chen K, Selitsky SR, et al. Identification of driver genes in hepatocellular carcinoma by exome sequencing. Hepatology 2013;58:1693-702.
5. Kan Z, Zheng H, Liu X, Li S, Barber TD, et al. Whole-genome sequencing identifies recurrent mutations in hepatocellular carcinoma. Genome Res 2013;23:1422-33.
7. Peng Y, Croce CM. The role of MicroRNAs in human cancer. Signal Transduct Target Ther 2016;1:15004.
8. Kim VN, Han J, Siomi MC. Biogenesis of small RNAs in animals. Nat Rev Mol Cell Biol 2009;10:126-39.
9. Cai X, Hagedorn CH, Cullen BR. Human microRNAs are processed from capped, polyadenylated transcripts that can also function as mRNAs. RNA 2004;10:1957-66.
10. Kosaka N, Iguchi H, Yoshioka Y, Takeshita F, Matsuki Y, et al. Secretory mechanisms and intercellular transfer of microRNAs in living cells. J Biol Chem 2010;285:17442-52.
11. Alberti C, Cochella L. A framework for understanding the roles of miRNAs in animal development. Development 2017;144:2548-59.
12. Andrés-León E, Cases I, Alonso S, Rojas AM. Novel miRNA-mRNA interactions conserved in essential cancer pathways. Sci Rep 2017;7:46101.
13. Wong CCL, Wong CM, Tung EKK, Au SLK, Lee JMF, et al. The microRNA miR-139 suppresses metastasis and progression of hepatocellular carcinoma by down-regulating Rho-kinase 2. Gastroenterology 2011;140:322-31.
14. Gu W, Li X, Wang J. miR-139 regulates the proliferation and invasion of hepatocellular carcinoma through the WNT/TCF-4 pathway. Oncol Rep 2014;31:397-404.
15. Nagy A, Lanczky A, Menyhart O, Gyorffy B. Validation of miRNA prognostic power in hepatocellular carcinoma using expression data of independent datasets. Sci Rep 2018;8:9227.
16. Li Y, Di C, Li W, Cai W, Tan X, et al. Oncomirs miRNA-221/222 and tumor suppressors miRNA-199a/195 are crucial miRNAs in liver cancer: a systematic analysis. Dig Dis Sci 2016;61:2315-27.
17. Mou T, Zhu D, Wei X, Li T, Zheng D, et al. Identification and interaction analysis of key genes and microRNAs in hepatocellular carcinoma by bioinformatics analysis. World J Surg Oncol 2017;15:63.
18. Shi KQ, Lin Z, Chen XJ, Song M, Wang YQ, et al. Hepatocellular carcinoma associated microRNA expression signature: integrated bioinformatics analysis, experimental validation and clinical significance. Oncotarget 2015;6:25093-108.
19. Lou W, Liu J, Ding B, Chen D, Xu L, et al. Identification of potential miRNA-mRNA regulatory network contributing to pathogenesis of HBV-related HCC. J Transl Med 2019;17:7.
20. Wang JL, Hu Y, Kong X, Wang ZH, Chen HY, et al. Candidate microRNA biomarkers in human gastric cancer: a systematic review and validation study. PLoS One 2013;8:e73683.
21. Moher D, Liberati A, Tetzlaff J, Altman DG, Group P. Preferred reporting items for systematic reviews and meta-analyses: the PRISMA statement. PLoS Med 2009;6:e1000097.
22. Witjes CDM, van Aalten SM, Steyerberg EW, Borsboom GJJM, de Man RA, et al. Recently introduced biomarkers for screening of hepatocellular carcinoma: a systematic review and meta-analysis. Hepatol Int 2013;7:59-64.
23. le Sage C, Nagel R, Egan DA, Schrier M, Mesman E, et al. Regulation of the p27(Kip1) tumor suppressor by miR-221 and miR-222 promotes cancer cell proliferation. EMBO J 2007;26:3699-708.
24. Fornari F, Gramantieri L, Ferracin M, Veronese A, Sabbioni S, et al. MiR-221 controls CDKN1C/p57 and CDKN1B/p27 expression in human hepatocellular carcinoma. Oncogene 2008;27:5651-61.
25. Santhekadur PK, Das SK, Gredler R, Chen D, Srivastava J, et al. Multifunction protein staphylococcal nuclease domain containing 1 (SND1) promotes tumor angiogenesis in human hepatocellular carcinoma through novel pathway that involves nuclear factor kappaB and miR-221. J Biol Chem 2012;287:13952-8.
27. Landgraf P, Rusu M, Sheridan R, Sewer A, Iovino N, et al. A mammalian microRNA expression atlas based on small RNA library sequencing. Cell 2007;129:1401-14.
28. Ying Q, Liang L, Guo W, Zha R, Tian Q, et al. Hypoxia-inducible miR-210 augments the metastatic potential of tumor cells by targeting vacuole membrane protein 1 in hepatocellular carcinoma. Hepatology 2011;54:2064-75.
29. Huang X, Ding L, Bennewith KL, Tong RT, Welford SM, et al. Hypoxia-inducible mir-210 regulates normoxic gene expression involved in tumor initiation. Mol Cell 2009;35:856-67.
30. Zhang Z, Sun H, Dai H, Walsh RM, Imakura M, et al. MicroRNA miR-210 modulates cellular response to hypoxia through the MYC antagonist MNT. Cell Cycle 2009;8:2756-68.
31. Chen PJ, Yeh SH, Liu WH, Lin CC, Huang HC, et al. Androgen pathway stimulates microRNA-216a transcription to suppress the tumor suppressor in lung cancer-1 gene in early hepatocarcinogenesis. Hepatology 2012;56:632-43.
32. Lan SH, Wu SY, Zuchini R, Lin XZ, Su IJ, et al. Autophagy-preferential degradation of MIR224 participates in hepatocellular carcinoma tumorigenesis. Autophagy 2014;10:1687-9.
33. Fornari F, Milazzo M, Chieco P, Negrini M, Calin GA, et al. MiR-199a-3p regulates mTOR and c-Met to influence the doxorubicin sensitivity of human hepatocarcinoma cells. Cancer Res 2010;70:5184-93.
34. Murakami Y, Aly HH, Tajima A, Inoue I, Shimotohno K. Regulation of the hepatitis C virus genome replication by miR-199a. J Hepatol 2009;50:453-60.
35. Hou J, Lin L, Zhou W, Wang Z, Ding G, et al. Identification of miRNomes in human liver and hepatocellular carcinoma reveals miR-199a/b-3p as therapeutic target for hepatocellular carcinoma. Cancer Cell 2011;19:232-43.
36. Guo W, Qiu Z, Wang Z, Wang Q, Tan N, et al. MiR-199a-5p is negatively associated with malignancies and regulates glycolysis and lactate production by targeting hexokinase 2 in liver cancer. Hepatology 2015;62:1132-44.
37. Furuta M, Kozaki K, Tanimoto K, Tanaka S, Arii S, et al. The tumor-suppressive miR-497-195 cluster targets multiple cell-cycle regulators in hepatocellular carcinoma. PloS One 2013;8:e60155.
38. Lagos-Quintana M, Rauhut R, Yalcin A, Meyer J, Lendeckel W, et al. Identification of tissue-specific microRNAs from mouse. Curr Biol 2002;12:735-9.
39. Girard M, Jacquemin E, Munnich A, Lyonnet S, Henrion-Caude A. miR-122, a paradigm for the role of microRNAs in the liver. J Hepatol 2008;48:648-56.
40. Hou W, Tian Q, Zheng J, Bonkovsky HL. MicroRNA-196 represses Bach1 protein and hepatitis C virus gene expression in human hepatoma cells expressing hepatitis C viral proteins. Hepatology 2010;51:1494-504.
41. Krützfeldt J, Rajewsky N, Braich R, Rajeev KG, Tuschl T, et al. Silencing of microRNAs in vivo with ‘antagomirs’. Nature 2005;438:685-9.
42. Esau C, Davis S, Murray SF, Yu XX, Pandey SK, et al. MiR-122 regulation of lipid metabolism revealed by in vivo antisense targeting. Cell Metab 2006;3:87-98.
43. Elmén J, Lindow M, Schütz S, Lawrence M, Petri A, et al. LNA-mediated microRNA silencing in non-human primates. Nature 2008;452:896-9.
44. Tsai WC, Hsu SD, Hsu CS, Lai TC, Chen SJ, et al. MicroRNA-122 plays a critical role in liver homeostasis and hepatocarcinogenesis. J Clin Invest 2012;122:2884-97.
45. Simerzin A, Zorde-Khvalevsky E, Rivkin M, Adar R, Zucman-Rossi J, et al. The liver-specific microRNA-122*, the complementary strand of microRNA-122, acts as a tumor suppressor by modulating the p53/mouse double minute 2 homolog circuitry. Hepatology 2016;64:1623-36.
46. Roy S, Hooiveld GJ, Seehawer M, Caruso S, Heinzmann F, et al. microRNA 193a-5p regulates levels of nucleolar- and spindle-associated protein 1 to suppress hepatocarcinogenesis. Gastroenterology 2018;155:1951-66.
47. Raemaekers T, Ribbeck K, Beaudouin J, Annaert W, Van Camp M, et al. NuSAP, a novel microtubule-associated protein involved in mitotic spindle organization. J Cell Biol 2003;162:1017-29.
48. Wu H, Tao J, Li X, Zhang T, Zhao L, et al. MicroRNA-206 prevents the pathogenesis of hepatocellular carcinoma via modulating expression of cMet and Cdk6. Hepatology 2017;66:1952-67.
49. Nejak-Bowen KN, Monga SP. Beta-catenin signaling, liver regeneration and hepatocellular cancer: sorting the good from the bad. Semin Cancer Biol 2011;21:44-58.
50. Zhang JG, Shi Y, Hong DF, Song M, Huang D, et al. MiR-148b suppresses cell proliferation and invasion in hepatocellular carcinoma by targeting WNT1/beta-catenin pathway. Sci Rep 2015;5:8087.
51. You Y, Que K, Zhou Y, Zhang Z, Zhao X, et al. MicroRNA-766-3p Inhibits tumour progression by targeting Wnt3a in hepatocellular carcinoma. Mol Cells 2018;41:830-41.
52. Meng F, Henson R, Wehbe-Janek H, Ghoshal K, Jacob ST, et al. MicroRNA-21 regulates expression of the PTEN tumor suppressor gene in human hepatocellular cancer. Gastroenterology 2007;133:647-58.
53. Wu H, Ng R, Chen X, Steer CJ, Song G. MicroRNA-21 is a potential link between non-alcoholic fatty liver disease and hepatocellular carcinoma via modulation of the HBP1-p53-Srebp1c pathway. Gut 2016;65:1850-60.
54. Rong M, Chen G, Dang Y. Increased miR-221 expression in hepatocellular carcinoma tissues and its role in enhancing cell growth and inhibiting apoptosis in vitro. BMC Cancer 2013;13:21.
55. Pineau P, Volinia S, McJunkin K, Marchio A, Battiston C, et al. miR-221 overexpression contributes to liver tumorigenesis. Proc Natl Acad Sci U S A 2010;107:264-9.
56. Liu Z, Sun J, Liu B, Zhao M, Xing E, et al. miRNA-222 promotes liver cancer cell proliferation, migration and invasion and inhibits apoptosis by targeting BBC3. Int J Mol Med 2018;42:141-8.
57. Ma D, Tao X, Gao F, Fan C, Wu D. miR-224 functions as an onco-miRNA in hepatocellular carcinoma cells by activating AKT signaling. Oncol Lett 2012;4:483-8.
58. Yu L, Zhang J, Guo X, Li Z, Zhang P. MicroRNA-224 upregulation and AKT activation synergistically predict poor prognosis in patients with hepatocellular carcinoma. Cancer Epidemiol 2014;38:408-13.
59. Wu GG, Li WH, He WG, Jiang N, Zhang GX, et al. Mir-184 post-transcriptionally regulates SOX7 expression and promotes cell proliferation in human hepatocellular carcinoma. PLoS One 2014;9:e88796.
60. Ohta K, Hoshino H, Wang J, Ono S, Iida Y, et al. MicroRNA-93 activates c-Met/PI3K/Akt pathway activity in hepatocellular carcinoma by directly inhibiting PTEN and CDKN1A. Oncotarget 2015;6:3211-24.
61. Sun C, Yao X, Jiang Q, Sun X. miR-106b targets DAB2 to promote hepatocellular carcinoma cell proliferation and metastasis. Oncol Lett 2018;16:3063-9.
62. Giovannini C, Minguzzi M, Baglioni M, Fornari F, Giannone F, et al. Suppression of p53 by Notch3 is mediated by Cyclin G1 and sustained by MDM2 and miR-221 axis in hepatocellular carcinoma. Oncotarget 2014;5:10607-20.
63. Ramassone A, Pagotto S, Veronese A, Visone R. Epigenetics and microRNAs in cancer. Int J Mol Sci 2018;19.
64. Nojima M, Matsui T, Tamori A, Kubo S, Shirabe K, et al. Global, cancer-specific microRNA cluster hypomethylation was functionally associated with the development of non-b non-c hepatocellular carcinoma. Mol Cancer 2016;15:31.
65. He XX, Kuang SZ, Liao JZ, Xu CR, Chang Y, et al. The regulation of microRNA expression by DNA methylation in hepatocellular carcinoma. Mol Biosyst 2015;11:532-9.
66. Konno M, Koseki J, Asai A, Yamagata A, Shimamura T, et al. Distinct methylation levels of mature microRNAs in gastrointestinal cancers. Nat Commun 2019;10:3888.
67. Braconi C, Patel T. MicroRNA expression profiling: a molecular tool for defining the phenotype of hepatocellular tumors. Hepatology 2008;47:1807-9.
68. Yi PS, Li JS. High expression of miR-21 is not a predictor of poor prognosis in all patients with hepatocellular carcinoma. Mol Clin Oncol 2018;8:733-9.
69. Zhou Y, Ren H, Dai B, Li J, Shang L, et al. Hepatocellular carcinoma-derived exosomal miRNA-21 contributes to tumor progression by converting hepatocyte stellate cells to cancer-associated fibroblasts. J Exp Clin Cancer Res 2018;37:324.
70. Shih YT, Wang MC, Zhou J, Peng HH, Lee DY, et al. Endothelial progenitors promote hepatocarcinoma intrahepatic metastasis through monocyte chemotactic protein-1 induction of microRNA-21. Gut 2015;64:1132-47.
71. Li Q, Ding C, Chen C, Zhang Z, Xiao H, et al. miR-224 promotion of cell migration and invasion by targeting Homeobox D 10 gene in human hepatocellular carcinoma. J Gastroenterol Hepatol 2014;29:835-42.
72. Kogure T, Lin WL, Yan IK, Braconi C, Patel T. Intercellular nanovesicle-mediated microRNA transfer: a mechanism of environmental modulation of hepatocellular cancer cell growth. Hepatology 2011;54:1237-48.
73. Wei JX, Lv LH, Wan YL, Cao Y, Li GL, et al. Vps4A functions as a tumor suppressor by regulating the secretion and uptake of exosomal microRNAs in human hepatoma cells. Hepatology 2015;61:1284-94.
74. Lin XJ, Fang JH, Yang XJ, Zhang C, Yuan YF, et al. Hepatocellular carcinoma cell-secreted Exosomal MicroRNA-210 promotes angiogenesis in vitro and in vivo. Mol Ther Nucleic Acids 2018;11:243-52.
75. Fang JH, Zhang ZJ, Shang LR, Luo YW, Lin YF, et al. Hepatoma cell-secreted exosomal microRNA-103 increases vascular permeability and promotes metastasis by targeting junction proteins. Hepatology 2018;68:1459-75.
76. Fang T, Lv H, Lv G, Li T, Wang C, et al. Tumor-derived exosomal miR-1247-3p induces cancer-associated fibroblast activation to foster lung metastasis of liver cancer. Nat Commun 2018;9:191.
77. Greening DW, Gopal SK, Xu R, Simpson RJ, Chen WS. Exosomes and their roles in immune regulation and cancer. Semin Cell Dev Biol 2015;40:72-81.
78. Chen L, Gibbons DL, Goswami S, Cortez MA, Ahn YH, et al. Metastasis is regulated via microRNA-200/ZEB1 axis control of tumour cell PD-L1 expression and intratumoral immunosuppression. Nat Commun 2014;5:5241.
79. Cortez MA, Ivan C, Valdecanas D, Wang X, Peltier HJ, et al. PDL1 Regulation by p53 via miR-34. J Natl Cancer Inst 2015;108.
80. Ye L, Zhang Q, Cheng Y, Chen X, Wang G, et al. Tumor-derived exosomal HMGB1 fosters hepatocellular carcinoma immune evasion by promoting TIM-1(+) regulatory B-cell expansion. J Immunother Cancer 2018;6:145.
81. Yin C, Han Q, Xu D, Zheng B, Zhao X, et al. SALL4-mediated upregulation of exosomal miR-146a-5p drives T-cell exhaustion by M2 tumor-associated macrophages in HCC. Oncoimmunology 2019;8:1601479.
82. Liu J, Fan L, Yu H, Zhang J, He Y, et al. Endoplasmic reticulum stress causes liver cancer cells to release exosomal mir-23a-3p and up-regulate programmed death ligand 1 expression in macrophages. Hepatology 2019;70:241-58.
83. Ji J, Wang H, Li Y, Zheng L, Yin Y, et al. Diagnostic evaluation of des-gamma-carboxy prothrombin versus α-fetoprotein for hepatitis B virus-related hepatocellular carcinoma in China: a large-scale, multicentre study. PLoS One 2016;11:e0153227.
84. Lawrie CH, Gal S, Dunlop HM, Pushkaran B, Liggins AP, et al. Detection of elevated levels of tumour-associated microRNAs in serum of patients with diffuse large B-cell lymphoma. Br J Haematol 2008;141:672-5.
85. Borel F, Konstantinova P, Jansen PL. Diagnostic and therapeutic potential of miRNA signatures in patients with hepatocellular carcinoma. J Hepatol 2012;56:1371-83.
86. Zhou J, Yu L, Gao X, Hu J, Wang J, et al. Plasma microRNA panel to diagnose hepatitis B virus- related hepatocellular carcinoma. J Clin Oncol 2011;29:4781-8.
87. Jin Y, Wong YS, Goh BKP, Chan CY, Cheow PC, et al. Circulating microRNAs as potential diagnostic and prognostic biomarkers in hepatocellular carcinoma. Sci Rep 2019;9:10464.
88. Tan Y, Ge G, Pan T, Wen D, Chen L, et al. A serum microRNA panel as potential biomarkers for hepatocellular carcinoma related with hepatitis B virus. PLoS One 2014;9:e107986.
89. Zhang Y, Li T, Qiu Y, Zhang T, Guo P, et al. Serum microRNA panel for early diagnosis of the onset of hepatocellular carcinoma. Medicine (Baltimore) 2017;96:e5642.
90. Huang JT, Liu SM, Ma H, Yang Y, Zhang X, et al. Systematic review and meta-analysis: circulating miRNAs for diagnosis of hepatocellular carcinoma. J Cell Physiol 2016;231:328-35.
91. Yamamoto Y, Kondo S, Matsuzaki J, Esaki M, Okusaka T, et al. Highly sensitive circulating microRNA panel for accurate detection of hepatocellular carcinoma in patients with liver disease. Hepatol Commun 2020;4:284-97.
92. Foye C, Yan IK, David W, Shukla N, Habboush Y, et al. Comparison of miRNA quantitation by Nanostring in serum and plasma samples. PLoS One 2017;12:e0189165.
93. Xu Y, Bu X, Dai C, Shang C. High serum microRNA-122 level is independently associated with higher overall survival rate in hepatocellular carcinoma patients. Tumour Biol 2015;36:4773-6.
94. Kim SS, Nam JS, Cho HJ, Won JH, Kim JW, et al. Plasma micoRNA-122 as a predictive marker for treatment response following transarterial chemoembolization in patients with hepatocellular carcinoma. J Gastroenterol Hepatol 2017;32:199-207.
95. Liu M, Liu J, Wang L, Wu H, Zhou C, et al. Association of serum microRNA expression in hepatocellular carcinomas treated with transarterial chemoembolization and patient survival. PloS One 2014;9:e109347.
96. Li J, Wang Y, Yu W, Chen J, Luo J. Expression of serum miR-221 in human hepatocellular carcinoma and its prognostic significance. Biochem Biophys Res Commun 2011;406:70-3.
97. Zhang Y, Wei C, Guo CC, Bi RX, Xie J, et al. Prognostic value of microRNAs in hepatocellular carcinoma: a meta-analysis. Oncotarget 2017;8:107237-57.
98. Shruthi K, Makol A, Anuradha C. Sorafenib response in hepatocellular carcinoma: microRNAs as tuning forks. Hepatol Res 2018;48:5-14.
99. Chuma M, Toyoda H, Matsuzaki J, Saito Y, Kumada T, et al. Circulating microRNA-1246 as a possible biomarker for early tumor recurrence of hepatocellular carcinoma. Hepatol Res 2019;49:810-22.
100. Cho HJ, Kim SS, Nam JS, Kim JK, Lee JH, et al. Low levels of circulating microRNA-26a/29a as poor prognostic markers in patients with hepatocellular carcinoma who underwent curative treatment. Clin Res Hepatol Gastroenterol 2017;41:181-9.
101. Nishida N, Arizumi T, Hagiwara S, Ida H, Sakurai T, et al. MicroRNAs for the prediction of early response to sorafenib treatment in human hepatocellular carcinoma. Liver Cancer 2017;6:113-25.
102. Fornari F, Pollutri D, Patrizi C, La Bella T, Marinelli S, et al. In hepatocellular carcinoma miR-221 modulates sorafenib resistance through inhibition of caspase-3-mediated apoptosis. Clin Cancer Res 2017;23:3953-65.
103. Teufel M, Seidel H, Köchert K, Meinhardt G, Finn RS, et al. Biomarkers associated with response to regorafenib in patients with hepatocellular carcinoma. Gastroenterology 2019;156:1731-41.
104. Chen X, Ba Y, Ma L, Cai X, Yin Y, et al. Characterization of microRNAs in serum: a novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res 2008;18:997-1006.
105. Melo SA, Luecke LB, Kahlert C, Fernandez AF, Gammon ST, et al. Glypican-1 identifies cancer exosomes and detects early pancreatic cancer. Nature 2015;523:177-82.
106. Pant S, Hilton H, Burczynski ME. The multifaceted exosome: biogenesis, role in normal and aberrant cellular function, and frontiers for pharmacological and biomarker opportunities. Biochem Pharmacol 2012;83:1484-94.
107. Pan JH, Zhou H, Zhao XX, Ding H, Li W, et al. Role of exosomes and exosomal microRNAs in hepatocellular carcinoma: Potential in diagnosis and antitumour treatments (Review). Int J Mol Med 2018;41:1809-16.
108. Sohn W, Kim J, Kang SH. Serum exosomal microRNAs as novel biomarkers for hepatocellular carcinoma. Exp Mol Med 2015;47:e184.
109. Lee YR, Kim G, Tak WY, Jang SY, Kweon YO, et al. Circulating exosomal non-coding RNAs as prognostic biomarkers in human hepatocellular carcinoma. Int J Cancer 2019;144:1444-52.
110. Afonso MB, Rodrigues PM, Simão AL, Castro RE. Circulating microRNAs as potential biomarkers in non-alcoholic fatty liver disease and hepatocellular carcinoma. J Clin Med 2016;5:30.
111. Liu W, Chen S, Liu B. Diagnostic and prognostic values of serum exosomal microRNA-21 in children with hepatoblastoma: a Chinese population-based study. Pediatr Surg Int 2016;32:1059-65.
112. Mjelle R, Dima SO, Bacalbasa N, Chawla K, Sorop A, et al. Comprehensive transcriptomic analyses of tissue, serum, and serum exosomes from hepatocellular carcinoma patients. BMC Cancer 2019;19:1007.
113. Sugimachi K, Matsumura T, Hirata H, Uchi R, Ueda M, et al. Identification of a bona fide microRNA biomarker in serum exosomes that predicts hepatocellular carcinoma recurrence after liver transplantation. Br J Cancer 2015;112:532-8.
114. Suehiro T, Miyaaki H, Kanda Y, Shibata H, Honda T, et al. Serum exosomal microRNA-122 and microRNA-21 as predictive biomarkers in transarterial chemoembolization-treated hepatocellular carcinoma patients. Oncol Lett 2018;16:3267-73.
115. Yuan JH, Yang F, Wang F, Ma JZ, Guo YJ, et al. A long noncoding RNA activated by TGF-β promotes the invasion-metastasis cascade in hepatocellular carcinoma. Cancer Cell 2014;25:666-81.
116. Hoshino A, Costa-Silva B, Shen TL, Rodrigues G, Hashimoto A, et al. Tumour exosome integrins determine organotropic metastasis. Nature 2015;527:329-35.
117. Ling H, Fabbri M, Calin GA. MicroRNAs and other non-coding RNAs as targets for anticancer drug development. Nat Rev Drug Discov 2013;12:847-65.
118. Fu Y, Chen J, Huang Z. Recent progress in microRNA-based delivery systems for the treatment of human disease. ExRNA 2019;1:24.
119. Lindow M, Kauppinen S. Discovering the first microrna-targeted drug. J Cell Biol 2012;199:407-12.
120. Gebert LF, Rebhan MA, Crivelli SE, Denzler R, Stoffel M, et al. Miravirsen (SPC3649) can inhibit the biogenesis of miR-122. Nucleic Acids Res 2014;42:609-21.
122. Jopling CL, Schütz S, Sarnow P. Position-dependent function for a tandem microRNA miR-122-binding site located in the hepatitis C virus RNA genome. Cell Host Microbe 2008;4:77-85.
123. Trajkovski M, Hausser J, Soutschek J, Bhat B, Akin A, et al. MicroRNAs 103 and 107 regulate insulin sensitivity. Nature 2011;474:649-53.
124. . Regulus Therapeutics I. RG-125 (AZD4076), a microRNA therapeutic targeting microRNA 103/107 for the treatment of NASH in patients with type 2 diabetes/Pre-Diabetes, selected as clinical candidate by AstraZeneca. Press release; 2015.
125. Aravalli RN. Development of microRNA therapeutics for hepatocellular carcinoma. Diagnostics (Basel) 2013;3:170-91.
126. Daige CL, Wiggins JF, Priddy L, Nelligan-Davis T, Zhao J, et al. Systemic delivery of a miR34a mimic as a potential therapeutic for liver cancer. Mol Cancer Ther 2014;13:2352-60.
127. Beg MS, Brenner AJ, Sachdev J, Borad M, Kang YK, et al. Phase I study of MRX34, a liposomal miR-34a mimic, administered twice weekly in patients with advanced solid tumors. Invest New Drugs 2017;35:180-8.
128. Sato Y, Hatakeyama H, Sakurai Y, Hyodo M, Akita H, et al. A pH-sensitive cationic lipid facilitates the delivery of liposomal siRNA and gene silencing activity in vitro and in vivo. J Control Release 2012;163:267-76.
129. Yamamoto N, Sato Y, Munakata T, Kakuni M, Tateno C, et al. Novel pH-sensitive multifunctional envelope-type nanodevice for siRNA-based treatments for chronic HBV infection. J Hepatol 2016;64:547-55.
130. Santangelo L, Battistelli C, Montaldo C, Citarella F, Strippoli R, et al. Functional roles and therapeutic applications of exosomes in hepatocellular carcinoma. Biomed Res Int 2017;2017:2931813.
131. Lötvall J, Hill AF, Hochberg F, Buzás EI, Di Vizio D, et al. Minimal experimental requirements for definition of extracellular vesicles and their functions: a position statement from the International Society for Extracellular Vesicles. J Extracell Vesicles 2014;3:26913.
132. Gould SJ, Raposo G. As we wait: coping with an imperfect nomenclature for extracellular vesicles. J Extracell Vesicles 2013;2.
133. Ingato D, Lee JU, Sim SJ, Kwon YJ. Good things come in small packages: Overcoming challenges to harness extracellular vesicles for therapeutic delivery. J Control Release 2016;241:174-85.
134. Pomatto MAC, Bussolati B, D’Antico S, Ghiotto S, Tetta C, et al. Improved loading of plasma-derived extracellular vesicles to encapsulate antitumor miRNAs. Mol Ther Methods Clin Dev 2019;13:133-44.
Cite This Article
Export citation file: BibTeX | RIS
OAE Style
Onishi M, Ochiya T, Tanaka Y. MicroRNA and liver cancer. Cancer Drug Resist 2020;3:385-400. http://dx.doi.org/10.20517/cdr.2019.110
AMA Style
Onishi M, Ochiya T, Tanaka Y. MicroRNA and liver cancer. Cancer Drug Resistance. 2020; 3(3): 385-400. http://dx.doi.org/10.20517/cdr.2019.110
Chicago/Turabian Style
Onishi, Masaya, Takahiro Ochiya, Yasuhito Tanaka. 2020. "MicroRNA and liver cancer" Cancer Drug Resistance. 3, no.3: 385-400. http://dx.doi.org/10.20517/cdr.2019.110
ACS Style
Onishi, M.; Ochiya T.; Tanaka Y. MicroRNA and liver cancer. Cancer Drug Resist. 2020, 3, 385-400. http://dx.doi.org/10.20517/cdr.2019.110
About This Article
Special Issue
Copyright
Data & Comments
Data
 Cite This Article 39 clicks
Cite This Article 39 clicks













Comments
Comments must be written in English. Spam, offensive content, impersonation, and private information will not be permitted. If any comment is reported and identified as inappropriate content by OAE staff, the comment will be removed without notice. If you have any queries or need any help, please contact us at support@oaepublish.com.